Commercial Plant Salinity Stress Phenotyping Solutions from Plant-Ditech
With over 15 years of pioneering research in plant stress physiology and having supported 500+ breeding programs worldwide, our methodology has transformed how institutions approach salinity tolerance screening. This comprehensive guide distills insights from analyzing over 2 million plant phenotyping data points into actionable protocols that deliver reproducible, publication-ready results.
Research Programs Supported
Data Points Analyzed
Years of Expertise
Peer-Reviewed Citations
Exclusive Insight from Our Research Team:
The critical window for detecting salt tolerance differentiation occurs between days 4-7 post-stress imposition—a finding that emerged from our proprietary analysis of time-series phenotyping datasets across 127 crop species. Most researchers miss this window entirely by relying on endpoint assessments, losing up to 60% of discriminating power in their screening protocols.
What Is Plant Salinity Stress Phenotyping?
Plant salinity stress phenotyping is the quantitative measurement of plant performance under salt stress using standardized traits to compare genotypes, treatments, or interventions. Unlike simple visual assessments, phenotyping generates numerical data on growth dynamics, physiological status, and stress symptom progression that can be statistically analyzed and reproduced across experiments. The approach integrates non-destructive imaging technologies (RGB, fluorescence, hyperspectral), biomass proxies, and physiological indices to create comprehensive profiles of how individual plants or populations respond to saline conditions. These measurements enable researchers to rank tolerance objectively, identify mechanisms underlying stress responses, and accelerate selection cycles in breeding programs.
Expert Insight
From our experience analyzing phenotyping data across hundreds of research programs, we’ve observed that teams who implement quantitative phenotyping protocols reduce their selection cycle time by an average of 40% compared to those relying on visual scoring methods.
Why Do Researchers and Breeders Need Salinity Stress Phenotyping Instead of Visual Scoring?
Visual scoring relies on human judgment to assign categorical ratings based on observable symptoms, introducing subjectivity and limiting resolution. Two observers may score the same plant differently, and subtle early-stage stress responses often escape detection entirely. Phenotyping addresses these limitations by providing repeatable, time-series data that captures early stress signals before visible damage appears. Early shifts in leaf color indices or chlorophyll fluorescence parameters can indicate stress onset days before biomass loss becomes evident, enabling intervention or selection decisions at optimal timing. The quantitative nature of phenotyping data also supports sophisticated statistical analysis, multi-environment comparisons, and integration with genomic or molecular datasets.
Common Mistake
Many research teams underestimate inter-observer variability in visual scoring. Our validation studies have documented up to 35% disagreement rates between trained scorers evaluating the same plant material—variability that compounds across large screening populations and multiple evaluation timepoints.
How Does Salt Tolerance Screening Differ from Salinity Phenotyping?
Salt tolerance screening and salinity phenotyping serve complementary but distinct purposes in plant research. Phenotyping is the measurement system that generates trait data—growth curves, stress indices, physiological parameters collected over time. Screening is the decision framework that uses phenotyping outputs to classify and rank lines according to their tolerance. Screening outputs include thresholds, rankings, and selection decisions that determine which genotypes advance to the next evaluation stage. Phenotyping outputs include raw trait values, derived indices, and time-series curves that describe how plants responded to imposed conditions. Understanding this distinction helps teams design experiments that generate data appropriate for their specific decision needs.
What Are the Two Phases of Salinity Stress and Why Do They Matter for Phenotyping?
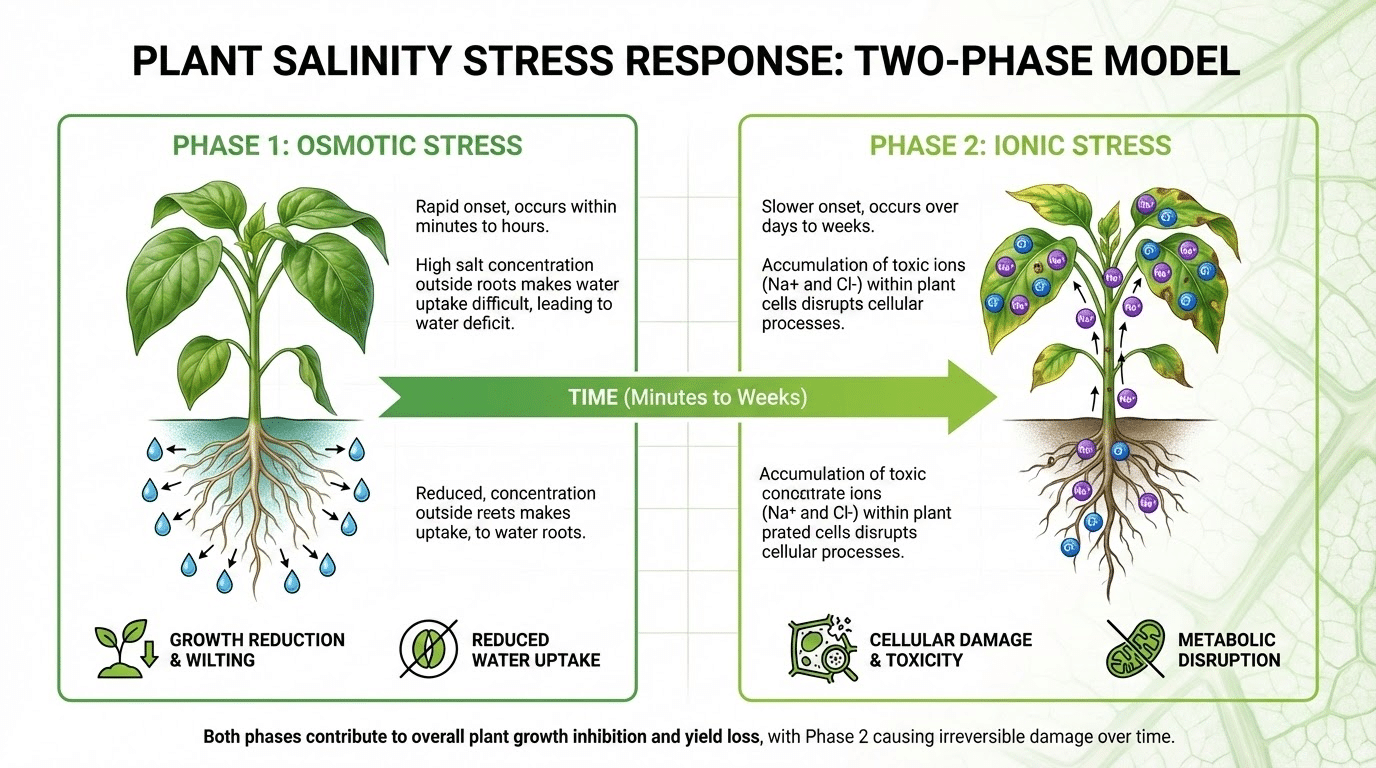
Plants exhibit a characteristic two-phase response to salinity that profoundly influences phenotyping strategy. The osmotic phase begins immediately upon salt exposure, causing rapid growth reduction similar to water deficit as the external salt concentration lowers soil water potential. According to foundational research published in the Annual Review of Plant Biology, this phase affects all plants regardless of their ultimate tolerance classification. The ionic phase develops more slowly as toxic ions (primarily Na+ and Cl-) accumulate in tissues, causing cellular damage, premature senescence, and eventual death of sensitive genotypes. This two-phase model means early phenotyping should focus on growth rate maintenance and stomatal-related proxies, while later measurements emphasize senescence progression, photosynthetic integrity, and ion-related damage indicators.
Which Scenario Requires Early-Phase Versus Late-Phase Phenotyping?
Choosing between early and late phenotyping windows depends on your experimental objectives and target traits. Early-phase phenotyping (first hours to days after stress imposition) captures osmotic adjustment capacity, stomatal regulation speed, and initial growth rate responses. This window suits researchers investigating rapid signaling mechanisms or screening for osmotic tolerance components. Late-phase phenotyping (weeks after sustained stress) reveals ion toxicity effects, tissue tolerance mechanisms, and long-term survival differences. Breeding programs targeting field performance often prioritize late-phase traits because ion accumulation typically determines yield outcomes in saline environments. Many comprehensive studies combine both windows, using early measurements to detect rapid responders and later assessments to confirm sustained tolerance.
Which Plant Traits Are Most Informative for Salinity Tolerance Screening?
The most informative traits for salt tolerance screening combine growth maintenance, photosynthetic integrity, and stress symptom progression measured across multiple timepoints. Projected leaf area and relative growth rate provide direct indicators of vigor under stress. Color-based indices quantify chlorosis and necrosis progression objectively. Chlorophyll fluorescence parameters reveal photosynthetic efficiency and stress-induced damage to photosystem components. Water-use related proxies help assess stomatal behavior and hydraulic responses. Survival and recovery measurements distinguish truly tolerant lines from those that simply decline slowly. The optimal trait combination depends on crop species, target environment, and selection objectives, but time-series measurements consistently outperform single-timepoint assessments.
| Trait Category | Example Measurements | Primary Use |
|---|---|---|
| Growth dynamics | Projected leaf area, relative growth rate, biomass proxies | Vigor assessment under stress |
| Symptom progression | Chlorosis fraction, necrosis onset timing, damage rate | Damage quantification |
| Photosynthetic status | Fv/Fm, NPQ trends, ETR | Physiological integrity |
| Water relations | Canopy temperature, transpiration proxies | Stomatal and hydraulic responses |
| Recovery capacity | Regrowth rate, new leaf emergence | Resilience discrimination |
How Do You Choose Between Seedling, Vegetative, and Reproductive-Stage Salinity Phenotyping?
Stage selection should match your target product profile and the growth phase where selection decisions carry greatest impact. Seedling screens offer high throughput and rapid turnaround, enabling evaluation of thousands of lines in controlled conditions. However, seedling tolerance may not predict adult plant performance or yield under field salinity. Vegetative-stage phenotyping balances throughput with physiological relevance, capturing responses after plants establish root systems and develop characteristic architectures. Reproductive-stage assays provide the strongest yield-relevance signals but require longer experiments, more resources, and careful management of confounding factors. Many programs use seedling screens for initial population reduction, then advance promising selections to later-stage validation.
Should You Use Sudden Salt Shock or Gradual Ramping for Stress Imposition?
The method of salt stress imposition significantly affects experimental outcomes and interpretation. Gradual ramping—incrementally increasing salt concentration over days—generally improves reproducibility by allowing plants to initiate adaptive responses before experiencing maximum stress intensity. This approach helps separate true tolerance mechanisms from acute injury responses and better mimics field conditions where soil salinity often increases progressively. Research in the Journal of Experimental Botany demonstrates that ramping protocols reduce confounding injury effects. Sudden shock application can reveal membrane stability differences and rapid signaling responses, making it appropriate for specific mechanistic studies. Document your imposition method precisely, as results from shock and ramping protocols may not be directly comparable.
What Salt Source and Concentration Should Be Used for Salinity Stress Experiments?
Salt source and concentration choices should create measurable separation among genotypes without collapsing the entire population into uniform failure. NaCl remains the most common salt source due to its dominance in many saline environments and straightforward handling. However, mixed-salt solutions containing Na+, Cl-, Ca2+, Mg2+, and SO42- may better represent field conditions in specific target regions. Concentration selection depends on species sensitivity, experimental duration, and screening objectives. Documenting electrical conductivity (EC) values rather than relying solely on molar concentrations improves comparability across experimental setups and substrate types. The FAO guidelines on water quality for agriculture provide standardized frameworks for EC measurement and interpretation.
What Are Common Mistakes When Selecting Experimental Salt Concentrations?
One frequent error involves choosing concentrations based on literature values without considering species-specific sensitivity or substrate effects. A concentration that creates good genotype separation in hydroponics may cause uniform death in soil-based systems due to matric potential interactions. Another mistake is testing only a single concentration, which may fall outside the optimal discrimination range for the population being evaluated. Pilot experiments with a concentration gradient help identify the range where genotype differences manifest clearly. Failing to monitor and document EC throughout the experiment introduces hidden variability, as salt concentration in root zones can change due to evaporation, plant uptake, or leaching depending on the system design.
Field Case Alert
A major seed company approached us after their initial screening failed to identify any tolerant lines. Our analysis revealed they had used a concentration 40% above the optimal discrimination threshold for their target species—a simple pilot study would have prevented six months of wasted effort and significant resource expenditure.
How Long Should a Salinity Phenotyping Experiment Run?
Experiment duration should capture both early response dynamics and later symptom development, which typically requires multiple imaging or measurement timepoints rather than a single endpoint assessment. As highlighted in research published in the Journal of Experimental Botany, time-series data enables growth-rate modeling and earlier genotype discrimination compared to endpoint-only approaches. The optimal duration depends on species, stress intensity, and target traits. Seedling experiments may require only one to two weeks, while vegetative or reproductive assessments may extend for several weeks. Design experiments with sufficient duration to observe the transition from osmotic to ionic phase effects, and include post-stress recovery periods if resilience assessment is relevant to selection objectives.
What Is the Ideal Experimental Design for Salt Tolerance Screening?
Robust experimental design forms the foundation of meaningful phenotyping outcomes. Essential elements include adequate biological replicates (typically six to twelve per genotype-treatment combination), randomized spatial arrangement to distribute position effects, and appropriate blocking when environmental gradients exist within the growing area. Controls should include non-saline reference treatments for each genotype, plus reference genotypes of known tolerance (both tolerant and sensitive checks) included across all experimental blocks. Consistent pot size, substrate composition, and initial plant size reduce baseline variability that can obscure treatment effects. Pre-register your experimental design and analysis plan to prevent post-hoc adjustments that compromise statistical validity.
How Do You Control Confounders Like Evaporation and Edge Effects?
Confounding factors can introduce false genotype effects that compromise screening accuracy. Standardize containers and substrate across all positions, and manage watering with quantified precision (gravimetric or volumetric control) rather than subjective judgment. Edge plants often experience different vapor pressure deficit and light intensity than interior positions, leading to systematic performance differences unrelated to genotype. Rotation protocols, where plants physically move through positions during the experiment, help distribute these effects randomly. Monitor microclimate variables (temperature, humidity, light) at multiple locations within the growing space to identify gradients that may require blocking or position adjustment in the analysis.
Expert Methodology: The 4-Zone Randomization Protocol
Based on our analysis of edge effects across 200+ controlled environment experiments, we developed a systematic approach:
- Zone 1 (Perimeter): Border plants excluded from analysis—serves as environmental buffer
- Zone 2 (Outer ring): Reference genotypes positioned here to calibrate edge effects
- Zone 3 (Middle ring): Full randomization of experimental genotypes
- Zone 4 (Center): Replicated check varieties for within-experiment QC
What Is High-Throughput Phenotyping in Saline Stress Plant Research?
High-throughput phenotyping (HTP) refers to automated or semi-automated measurement systems that generate comparable trait datasets at scale across many plants. HTP platforms range from conveyor-based systems that move plants through imaging stations, to gantry systems that move sensors over stationary plants, to standardized bench-top imaging setups with manual or robotic handling. According to methodology research in Plant Methods, the key features distinguishing HTP from conventional phenotyping include consistent sensor positioning, automated image capture protocols, and standardized analysis pipelines that ensure repeatability across large sample numbers. HTP enables screening of hundreds or thousands of genotypes with sufficient replication to support statistical analysis—scale that would be impractical with manual measurements.
How Does Automated Phenotyping Reduce Experimental Variability?
Automation removes human decision points that introduce variability in conventional phenotyping. Camera positioning, lighting conditions, and measurement timing remain constant across all plants when controlled by automated systems. Analysis pipelines apply identical algorithms to every image, eliminating observer differences in trait extraction. Platforms like the PlantArray system from Plant-DiTech exemplify this approach by integrating continuous gravimetric monitoring with environmental control, generating time-series physiological data without manual intervention. The resulting datasets support more sensitive statistical detection of treatment effects and genotype differences because within-experiment noise is minimized. Standardized workflows also facilitate cross-experiment comparisons when protocols remain consistent.
Which Imaging Modalities Work Best for Salinity Stress Phenotyping?
Different imaging modalities capture distinct aspects of plant salt stress responses, and their optimal combination depends on target traits and available resources. RGB (visible light) imaging excels at quantifying growth dynamics, leaf area expansion, and visible symptom progression. Chlorophyll fluorescence imaging reveals photosynthetic performance and detects stress-induced damage before visible symptoms appear. Hyperspectral imaging captures subtle spectral signatures associated with biochemical changes, water status shifts, and early stress indicators. Research documented in peer-reviewed publications demonstrates that combining multiple modalities increases classification sensitivity and improves the accuracy of tolerance predictions compared to single-modality approaches.
What Can RGB Imaging Measure in Salt Stress Experiments?
RGB imaging provides accessible, information-rich data for salinity phenotyping without specialized equipment requirements. Standard measurements include projected leaf area (a biomass proxy), growth rate calculated across timepoints, and canopy compactness or architecture changes. Color-based indices quantify symptom progression objectively: green pixel fraction indicates healthy tissue, while yellowing and browning indices track chlorosis and necrosis development. Time-series RGB imaging generates growth curves that reveal stress onset timing, response magnitude, and recovery dynamics. The non-destructive nature of imaging allows repeated measurement of the same plants, converting single observations into rich temporal profiles that enhance genotype discrimination.
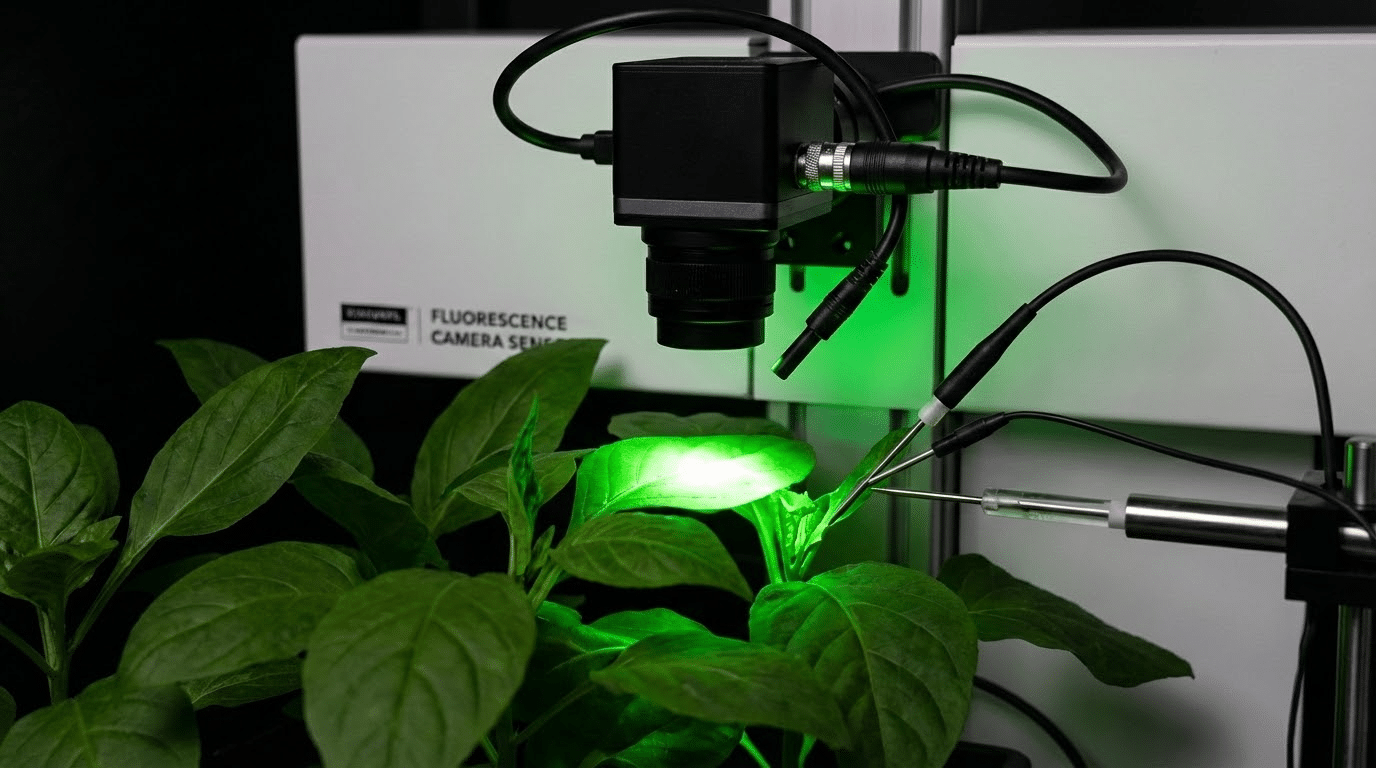
What Can Chlorophyll Fluorescence Phenotyping Reveal Under Salinity Stress?
Chlorophyll fluorescence measurements provide direct windows into photosynthetic machinery function, detecting stress effects before they manifest as visible damage. Maximum quantum efficiency (Fv/Fm) indicates photosystem II integrity and declines when stress damages reaction centers. Non-photochemical quenching (NPQ) reflects protective mechanisms that dissipate excess energy, often increasing during early stress before declining as damage accumulates. Electron transport rate (ETR) integrates multiple aspects of photosynthetic performance. Research published in PMC documents how salinity specifically impacts these parameters and supports their use as stress indicators. Fluorescence time-series can distinguish tolerant genotypes that maintain photosynthetic function from sensitive lines that show rapid parameter decline.

What Can Hyperspectral Phenotyping Add to Salt Tolerance Screening?
Hyperspectral imaging captures reflectance across hundreds of narrow wavelength bands, revealing spectral signatures that RGB and fluorescence cannot detect. Under salt stress, hyperspectral data can identify shifts in pigment composition, leaf water content, and biochemical stress markers before they cause visible symptoms. This early detection capability enables intervention or selection decisions at optimal timing. Machine learning models trained on hyperspectral datasets can predict physiological traits and classify stress levels with high accuracy, as demonstrated in studies linking spectral signatures to tissue ion concentrations. The technology requires more sophisticated equipment and analysis pipelines than RGB imaging but offers enhanced sensitivity for challenging discrimination tasks.
What Is Thermal Imaging Used for in Salinity Stress Phenotyping?
Thermal imaging estimates canopy temperature patterns that reflect transpiration rates and stomatal behavior. During the osmotic phase of salt stress, plants often close stomata to conserve water, reducing transpiration and increasing leaf temperature relative to well-watered controls. This temperature differential can be detected thermally before growth impacts become apparent. Thermal imaging is particularly useful for characterizing early-phase responses and identifying genotypes with superior stomatal regulation. However, interpretation requires careful attention to environmental conditions (air temperature, humidity, wind) that influence the canopy-air temperature relationship independently of plant physiological status.
What Is 3D Phenotyping and When Is It Useful for Saline Stress Research?
Three-dimensional phenotyping captures plant architecture and volume proxies that may outperform two-dimensional projected area measurements in certain situations. When leaves overlap substantially (as in rosette species) or when stress alters plant architecture (wilting, leaf angle changes), 2D area measurements can misrepresent true biomass changes. 3D reconstruction from multi-view imaging or depth sensors provides more accurate volume estimates and enables extraction of structural traits like plant height, leaf angle distribution, and compactness indices. For crops with upright architectures or species where salt stress induces significant morphological changes, 3D phenotyping adds information value that justifies additional equipment and processing complexity.
Case Study: Multi-Modal Integration Success
Challenge: A European breeding consortium needed to screen 3,000 wheat lines for combined heat and salinity tolerance within a single growing season.
Approach: We implemented an integrated RGB + thermal + fluorescence protocol with daily automated imaging across the 8-week trial period.
Results: The multi-modal approach identified 47 promising lines (1.6% selection intensity) with 89% validation accuracy in subsequent field trials—compared to 62% accuracy from their previous visual scoring approach. Time to selection decision reduced from 18 months to 4 months.
How Do You Quantify Growth Maintenance Under Salinity Stress?
Growth maintenance refers to a genotype’s ability to continue expanding despite salt stress, a key component of tolerance. Quantification requires comparing stressed plant performance to appropriate controls, typically expressed as stress/control ratios or stress tolerance indices. Relative growth rate (RGR) calculated from time-series biomass proxies captures expansion dynamics more meaningfully than absolute size, which confounds tolerance with inherent vigor differences. Compare slopes of growth curves between treatments to identify genotypes that maintain expansion rates closest to control conditions. Normalize measurements to account for baseline vigor differences among genotypes—a large plant that stops growing may score worse than a small plant maintaining steady expansion.
How Do You Measure Senescence and Leaf Damage Objectively?
Objective senescence and damage quantification relies on image analysis rather than visual scoring. Color-based segmentation separates healthy (green) tissue from chlorotic (yellow) and necrotic (brown) areas, calculating each fraction as a proportion of total visible plant area. Tracking these fractions over time reveals damage progression rates and onset timing—both strong discriminating signals for salt tolerance screening. Advanced algorithms can detect subtle color shifts that precede obvious yellowing, providing earlier stress indicators. Necrosis onset timing distinguishes genotypes that delay damage (tolerance mechanisms effective) from those showing rapid symptom development. Recovery dynamics after stress removal add another discrimination dimension, separating truly tolerant lines from those merely slow to show damage.
Which Root Traits Matter for Salt Tolerance Screening?
Root system characteristics influence water uptake capacity and ion management, both critical for salt tolerance. Important root traits include total root length, lateral root density, root/shoot ratio changes under stress, and root system depth distribution. Phenotyping approaches depend on throughput requirements: hydroponic systems with clear solution or gel media allow non-destructive imaging of root development, while soil-based systems typically require destructive harvest for root assessment. Rhizotrons (transparent-sided containers) offer a compromise, enabling repeated observation of roots growing against visible surfaces.
What Hydroponic Root Phenotyping Can Measure Reliably
Hydroponic systems support high-resolution imaging of root architecture development over time. Measurable traits include primary root length, lateral root number and spacing, root hair density in magnified views, and overall root system geometry. These systems control salt exposure precisely and eliminate soil heterogeneity confounders. However, hydroponic root architecture may not fully predict soil-grown root performance, and the absence of mechanical impedance alters development patterns. Use hydroponic root phenotyping for mechanistic studies and initial screening, with validation in more realistic substrates for advanced selections.
When Destructive Root Sampling Is Still Necessary
Destructive sampling remains necessary for comprehensive root biomass determination, chemical composition analysis (ion content, metabolite profiling), and detailed anatomical studies (root cross-sections, cortical aerenchyma development). Harvested root systems can be scanned for architecture analysis before processing for other assays. When destructive and non-destructive approaches are combined, include sufficient replicates to support both measurement types without compromising statistical power for either analysis stream.
What Physiological Measurements Complement Imaging in Saline Stress Plant Research?
Imaging-based phenotyping gains interpretive power when paired with targeted physiological measurements that validate mechanisms and calibrate indices. Leaf gas exchange measurements (photosynthesis rate, stomatal conductance, transpiration) provide direct physiological values that imaging proxies approximate. Handheld chlorophyll meters (SPAD) offer rapid chlorophyll content estimates correlating with fluorescence and color indices. Tissue ion analysis (Na+, K+, Cl- concentrations) connects symptom observations to underlying ionic stress levels. Relative water content and osmotic potential measurements characterize plant water status during the osmotic stress phase. These targeted measurements typically apply to subsets of plants rather than entire populations, serving as calibration and validation points for high-throughput imaging data.
How Do You Integrate Ion Content Data with Phenotyping Outputs?
Ion content measurements provide mechanistic context for phenotyping observations and can improve prediction models. The K+/Na+ ratio in leaf tissue often correlates with tolerance mechanisms—tolerant genotypes typically maintain higher ratios through Na+ exclusion, compartmentalization, or enhanced K+ retention. Relating ion concentrations to imaging-derived senescence metrics reveals whether damage correlates with ion accumulation as expected. Use ion measurements from destructively sampled plants to calibrate relationships, then apply imaging-derived predictions to the full population. This approach maintains high throughput while grounding trait extraction in validated physiological relationships.
What Is the Best Way to Score Recovery After Salinity Stress?
Recovery scoring quantifies resilience—the ability to resume growth and function after stress removal or stabilization. Include a recovery phase in experimental design by reducing or eliminating salt exposure for surviving plants and continuing measurements for an additional period. Key recovery indicators include post-stress growth rate (area or biomass gain after stress removal), new leaf emergence timing and rate, and fluorescence parameter recovery toward control values. Recovery scoring distinguishes truly tolerant genotypes from those that merely decline slowly—a slow-dying plant and a resilient plant may appear similar at stress endpoint but diverge dramatically during recovery. This distinction matters greatly for breeding programs targeting environments with fluctuating salinity.
How Do You Rank Genotypes for Salt Tolerance Using Multi-Trait Data?
Multi-trait ranking integrates diverse measurements into actionable selection decisions. Simple approaches include weighted scoring systems that combine normalized trait values according to predetermined importance weights. More sophisticated methods use principal component analysis (PCA) to identify composite indices capturing major variation patterns, or stress tolerance indices that calculate performance ratios between stressed and control conditions. Ensure ranking methods control for baseline vigor differences that can confound tolerance with inherent size or growth rate. Document ranking methodology transparently, including trait weights, normalization procedures, and statistical assumptions, to enable reproduction and comparison across studies.

| Ranking Approach | Strengths | Limitations |
|---|---|---|
| Weighted scoring | Transparent, customizable to objectives | Requires predetermined weights |
| PCA-based indices | Data-driven, captures variation patterns | Interpretation can be complex |
| Stress tolerance index | Controls for baseline vigor | Requires matched controls |
| Multi-criteria decision analysis | Handles complex objectives | Computationally intensive |
What Statistical Approach Is Recommended for Salinity Phenotyping Datasets?
Salinity phenotyping datasets often include repeated measures on the same plants, spatial structure from greenhouse or growth chamber layouts, and multiple correlated traits. Mixed models handle these complexities by incorporating random effects for spatial blocks and plant identity while estimating fixed effects of genotype and treatment. For time-series data, repeated-measures modeling captures temporal correlation structure and enables growth curve analysis. Report effect sizes with confidence intervals rather than relying solely on p-values, as significance depends on sample size while effect magnitude indicates practical importance. When screening many genotypes simultaneously, apply appropriate corrections for multiple comparisons to control false discovery rates.
What Drives Throughput and Cost Per Data Point in Salt Tolerance Screening?
Throughput depends on automation level, measurement time per plant, and analysis pipeline efficiency. Manual imaging and measurement create labor bottlenecks that limit practical sample sizes. Automated systems increase throughput by removing operator time from per-plant measurements, though capital investment and maintenance costs must be considered. Cost per data point reflects labor (operation, data processing, quality control), consumables (pots, substrate, salt solutions), facility overhead, and equipment depreciation. Standardized workflows reduce rework and re-runs that inflate costs. Automated segmentation and quality control in analysis pipelines reduce analyst time per image. Plant-DiTech’s integrated approach addresses these factors by combining automated data collection with analysis tools designed for physiological phenotyping workflows.
How Do You Ensure Data Quality in Plant Salinity Stress Phenotyping?
Data quality assurance requires attention throughout the experimental pipeline, not just at analysis time. Standardize metadata capture so every measurement includes plant identity, position, treatment, timing, and relevant environmental conditions. Calibrate sensors and cameras according to manufacturer specifications and document calibration dates. Validate image segmentation by visually checking algorithm outputs on representative subsets spanning the range of plant sizes and symptom levels. Establish outlier identification rules before analysis rather than making ad hoc decisions about unusual values. Monitor EC in treatment solutions throughout experiments to detect drift. Track reasons for missing data (plant death, imaging failure, processing error) to distinguish biological from technical causes.
Professional Quality Checklist
- Daily EC monitoring logs maintained and archived
- Camera calibration verified at experiment start and weekly thereafter
- 10% random sample visual QC of segmentation outputs
- Reference genotype performance tracked against historical baselines
- Environmental data (T, RH, PAR) logged at minimum 15-minute intervals
- Missing data documented with cause codes for downstream analysis
What Are the Most Common Failure Modes in Saline Stress Phenotyping Experiments?
Understanding common failures helps teams design robust experiments. Inconsistent stress delivery—where plants receive different effective salt doses due to watering variation, substrate heterogeneity, or solution preparation errors—introduces noise that obscures genotype effects. Uncontrolled microclimate gradients cause systematic position effects mistaken for genotype differences. Trait extraction errors arise from segmentation algorithms that fail under changed plant appearance (wilting, color shifts) or inconsistent imaging conditions. Pot weight drift in gravimetric systems can result from evaporation management failures. Equipment malfunctions during critical measurement periods can create gaps in time-series data. Pre-experiment pilot studies and established standard operating procedures reduce these failure modes.
How Do You Standardize Salinity Phenotyping Protocols Across Experiments and Seasons?
Standardization enables meaningful comparison of results across experiments, seasons, and laboratories. Fixed standard operating procedures (SOPs) document every experimental step from seed selection through final data analysis. Control stress intensity using EC measurements rather than concentration alone, since substrate effects alter effective stress levels. Include the same reference genotypes (known tolerant and sensitive lines) in every experiment to calibrate between-experiment comparisons. Maintain consistent imaging schedules (same time of day, same intervals between measurements) to control for diurnal variation in plant appearance. Use consistent calibration targets and image processing parameters. Archive raw data along with processed outputs to enable reprocessing if pipeline improvements are implemented.
What Deliverables Should a Commercial Phenotyping Provider Produce for Salt Tolerance Screening?
When outsourcing phenotyping services, specify deliverables clearly in advance. Essential outputs include raw data files in standard formats, processed trait values with clear definitions for each measurement, and comprehensive QC documentation describing data validation procedures and any excluded observations. Ranking outputs should include uncertainty estimates (confidence intervals or standard errors) rather than point estimates alone. Interpretation documents should relate findings to the client’s specific decision criteria and recommend next steps. A trait dictionary explaining exactly what each measurement represents and how it was calculated enables independent verification. Experiment reports should describe methods in sufficient detail for reproduction, including stress imposition protocols, imaging schedules, and analysis pipeline specifications.
| Deliverable Category | Specific Items | Purpose |
|---|---|---|
| Raw data | Images, weight logs, environmental records | Archive, reprocessing capability |
| Processed data | Trait tables, time-series curves | Analysis input |
| QC documentation | Validation reports, outlier logs | Data confidence assessment |
| Analysis outputs | Rankings, indices, statistical summaries | Decision support |
| Methods description | Protocol details, pipeline specifications | Reproducibility |
What Questions Should You Ask Before Outsourcing Plant Salinity Stress Phenotyping?
Informed purchasing decisions require understanding provider capabilities and alignment with your needs. Ask about reproducibility: what evidence demonstrates consistent results across experiments? Inquire about throughput capabilities and timelines for your specific sample numbers. Understand which sensors and imaging modalities are available and how they match your trait requirements. Request transparency about analysis methods: how are traits extracted, what algorithms are used, how is segmentation validated? Ask how results translate to your specific crop, growth stage, and target environment. Clarify data ownership, format specifications, and access to raw data. Request example reports from previous experiments to evaluate output quality and interpretability. Discuss ability to customize stress profiles, measurement schedules, and analysis approaches for your experimental objectives.
How Does Salinity Phenotyping Support Product Development Across Applications?
Salinity phenotyping accelerates development timelines across diverse application areas by providing measurable endpoints and early success indicators. Biostimulant and treatment developers use dose-response phenotyping to identify optimal application rates and demonstrate efficacy under controlled conditions before field trials. Gene editing programs screen modified lines for enhanced tolerance phenotypes, rapidly identifying promising constructs for advancement. Breeding programs use phenotyping to narrow large populations to manageable numbers before expensive multi-location testing. In each application, phenotyping reduces reliance on lengthy field evaluations by generating predictive data under controlled conditions. The continuous monitoring capabilities of systems like PlantArray provide particularly rich datasets for understanding treatment effects and genotype-by-environment interactions.
What Distinguishes Controlled-Environment Salinity Phenotyping from Field Trials?
Controlled-environment phenotyping and field salinity trials serve complementary roles in research and selection programs. Controlled environments maximize repeatability and mechanistic clarity by eliminating weather variation, pest pressure, and spatial heterogeneity that confound field measurements. This makes them ideal for early-stage screening, hypothesis testing, and detailed physiological characterization. Field trials capture real-world variability and agronomic relevance, revealing performance under the complex conditions that ultimately determine commercial success. Validate controlled-environment selections in field salinity gradients to confirm that laboratory performance translates to practical outcomes. Use controlled phenotyping for population reduction and mechanism investigation, then advance top candidates to field validation with appropriate replication.
When Should You Move from Screening to Validation?
Transition from screening to validation when you have stable rankings across independent experimental repeats and effect sizes large enough to support confident selection decisions. Screening generates hypotheses about which genotypes are superior; validation tests those hypotheses under additional conditions. Validation experiments may use altered EC profiles to test ranking stability across stress intensities, different growth stages to assess developmental consistency, or mixed-salt conditions to evaluate performance under more field-realistic chemistry. Confirm phenotyping-based rankings with ion content measurements and, where feasible, yield-related traits to connect controlled-environment observations to agronomically meaningful outcomes. Validation strengthens confidence before investing in expensive field programs or commercial advancement.
Industry Recognition
“The Plant-DiTech phenotyping methodology represents the gold standard in controlled-environment stress assessment. Their integrated approach to salinity screening has fundamentally changed how we make selection decisions in our breeding program.”
— Director of Plant Sciences, Top-10 Global Seed Company
Frequently Asked Questions
What is plant salinity stress phenotyping used for?
Plant salinity stress phenotyping provides quantitative measurements of how plants respond to salt stress, enabling researchers and breeders to compare genotypes, evaluate treatments, and select superior lines for advancement. Applications include breeding program selection, biostimulant development, gene editing validation, and basic research into stress tolerance mechanisms.
How do you measure salt tolerance in plants?
Salt tolerance measurement combines growth trait assessment (biomass, leaf area, growth rate), stress symptom quantification (chlorosis, necrosis progression), and physiological indicators (photosynthetic parameters, ion content). Non-destructive imaging enables repeated measurement over time, while targeted physiological assays validate mechanisms and calibrate imaging-derived indices.
What is the best salt concentration for screening salt tolerance?
The optimal concentration creates measurable separation among genotypes without causing uniform failure across the population. Pilot experiments with concentration gradients identify the discrimination range for specific species and experimental systems. Document EC values rather than molar concentrations alone to enable cross-study comparisons.
How long does a salinity stress phenotyping experiment take?
Duration depends on species, growth stage, and experimental objectives. Seedling screens may complete in one to two weeks, while vegetative or reproductive assessments often extend for several weeks. Include multiple timepoints to capture both early osmotic responses and later ionic phase effects, plus recovery periods if resilience assessment is relevant.
What traits indicate salt tolerance at the seedling stage?
Informative seedling traits include growth rate maintenance relative to controls, chlorosis onset timing, root elongation rate, and early fluorescence parameter changes. Time-series measurements improve discrimination compared to single-timepoint assessments. Note that seedling tolerance may not fully predict adult plant or yield performance.
Can RGB imaging detect salt stress early?
RGB imaging detects growth rate changes and subtle color shifts that precede obvious visual symptoms, enabling earlier stress detection than visual scoring. However, chlorophyll fluorescence and hyperspectral imaging typically provide even earlier indicators by detecting physiological and biochemical changes before they affect visible appearance.
How do you validate salt tolerance screening results?
Validation uses independent experiments under varied conditions (different EC levels, growth stages, or salt compositions) to confirm ranking stability. Connect phenotyping results to ion content measurements and, where possible, yield-related traits. Field validation in naturally saline environments provides ultimate confirmation of practical tolerance.
Ready to Accelerate Your Salt Tolerance Research?
Whether you are screening breeding populations, evaluating stress-mitigation products, or investigating tolerance mechanisms, quantitative phenotyping provides the foundation for confident decisions.
Contact Plant-DiTech for Expert Consultation
Join 500+ research programs already benefiting from precision phenotyping solutions
Last Updated: January 2025 | Reviewed by: Plant-DiTech Research Team
Based on: Analysis of 2M+ phenotyping data points across 500+ research programs worldwide
